Molecular Tools-3:
Modifying Enzymes:
In recombinant DNA technology a substantial number of enzymes are required for modifying DNA for a variety purposes. To modify the DNA, one should know what enzymes are required and their properties. It is sine quo non for any student of molecular Biotechnology.
Polymerases:
E. coli DNA pol-I:
Its Mol.wt is 109KD. This is extensively used for Nick translation and Random primer mode of labeling. It is also used in the second strand synthesis during cDNA preparation. However, it requires a primer or nick for its activity.
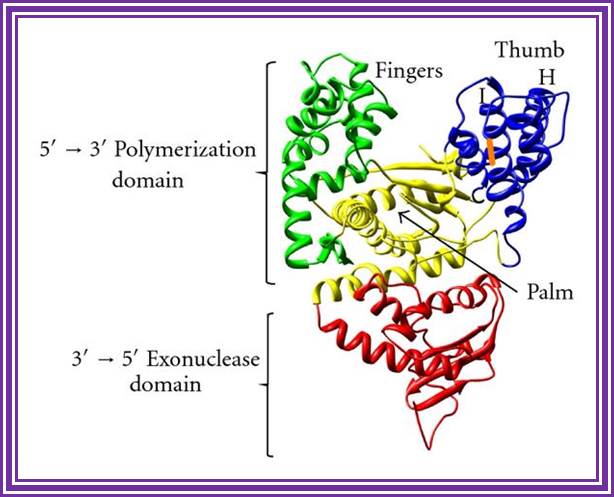
DNA Polymerase I; http://www.hindawi.com/
The enzyme from its NH3 end has 5’>3’ exonuclease, 3’>5’ exonuclease and 5’> 3’ palm with fingers has Polymerase activity. However the recombinant DNA pol-I is available as a truncated form called Klenow fragment (68KD), which has 3’>5’ exonuclease and 5’>>3’ Pol activity from its NH3 end. One has to be careful about this enzyme for; the concentration of dNTPs provided should be optimal otherwise instead of polymerase it acts as an exonuclease. If carefully used, one can apply this enzyme, for end filling or blunting dsDNA. Using primers one can produce ds DNA by what is called primer extension method. However, this enzyme has medium processivity and fidelity characters.
T7 DNA polymerase:
Its mol. Wt. is 112KD (87KD Pol + 12 KD theoredoxin 1:1). Using anti-theoredoxin antibody columns this enzyme can be purified to homogeneity. It is a viral enzyme. The virus infects E. coli and grows inside the cell. This polymerase is coded for by the viral genome and the host bacterial cell provides the theoredoxin.

http://www.personal.psu.edu/
Together the enzyme is excellent in its polymerase activity. It has 5’>>3’ polymerase activity and 3’>>5’ exonucleases activity. Recombinant methods have produced T7 pol without 3’>>5’ exonucleases property. This enzyme is called ‘Sequenase’ for this is an excellent enzyme for sequencing by primer extension protocols and even this can be used for labeling methods. This has high processivity and high-fidelity properties.
T4 DNA polymerase:
It is produced by the T4 Phage viral genome. Now it is available as recombinant product. Its Mol.wt is 114KD and consists of single polypeptide chain. This is used for DNA end filling and end labeling protocols. It has 5’>>3’ pol activity, 3’>>5’ exonucleases activity. The enzyme is quite-stable and it can be substituted for DNA pol-I. If there are any 3’ over hangs in dsDNA it can be removed by 3’ exonuclease activity and thus it produces blunt ends. On the contrary if the ds have 3’ recessive ends, the enzyme extends the 3’ ends, by 5 > 3’ polymerase activities. However, it lacks 5’> 3’ exonuclease activity.
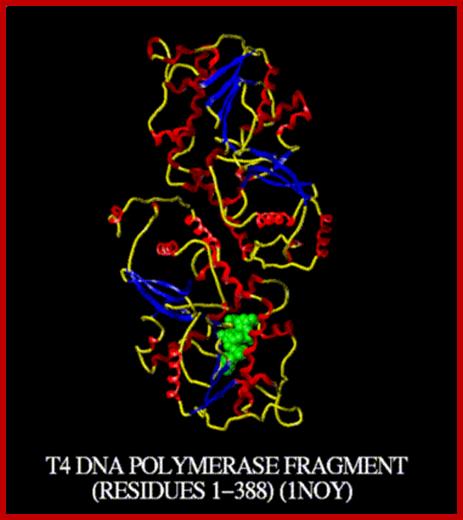
Biomolecules –-http://www.abren.net/image
Taq Polymerase:
This enzyme is isolated from thermophilic microbes, such as Thermophilous aquaticus, (Pyrococcus species), and Thermococcus litoralis. The enzyme Mol.wt is 85-95KD. They have apparent 5’>>3’ polymerase activity. And they don’t exhibit 3’>>5’ exonucleases activity. The enzyme is stable for 60 -70 minutes at 92^0 temperature. This enzyme is exclusively used for PCR amplification of DNA and it has low processivity and low fidelity characters. But recently pFu and ULTMA from Perkin Elmer/ Stratagene have been found to show high fidelity. These enzymes are exclusively and extensively used in PCR based florescent-labeled sequencing reactions.
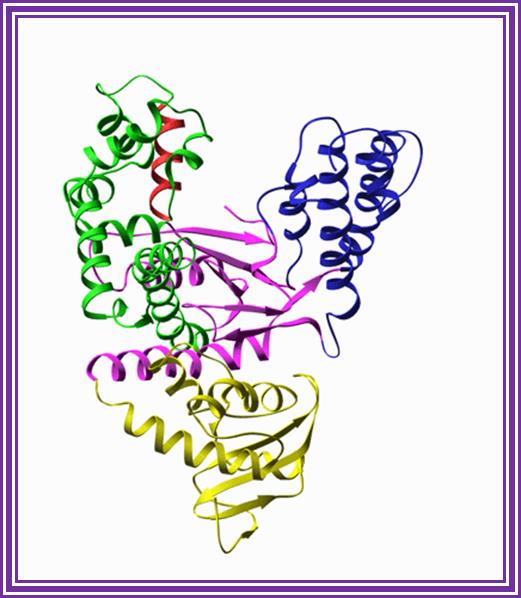
Taq polymerase:http://www.sharepoint.csiat,jmu.edu
These enzymes can also be used in Rt-PCR methods. These are also used in pathogen detection protocols. Also used in cDNA library preparations. The amplified ds strands have one A base as extensions. With suitable vectors the CDNA prepared can be directly ligated to dephosphorylated Vector DNAs.
RNA dependent DNA polymerase:
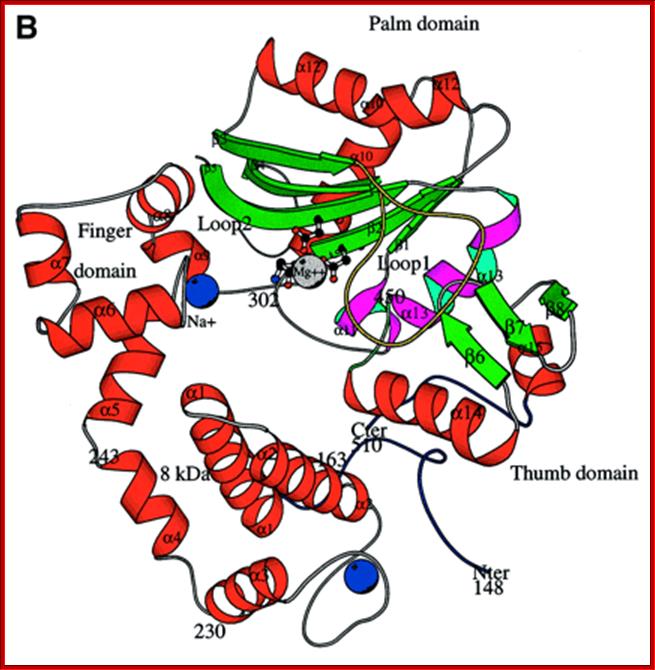
This is also called as Reverse Transcriptase. Its Mol.wt ranges from 85KD (from SDS-PAGE it shows 71KD) single polypeptide (murine melony leukemia virus) and 92 KD (two polypeptides called alpha 68KD and Beta 92KD) from Avian Myeloblastasis virus, all of them are retroviruses. These enzymes have RNA dependent DNA polymerase activity, but does not show any apparent 3’>>5’ exonucleases activity. It can also perform DNA dependent DNA polymerase activity. Another great feature of these enzymes is that they have RNase-H activity, where the enzyme removes by nicking or displacement of RNA when it is hybridized to DNA. It can remove nucleotides from both ends i.e. from 5’ and 3’ ends. The alpha subunit has contains both Polymerase and RNase -H activity.
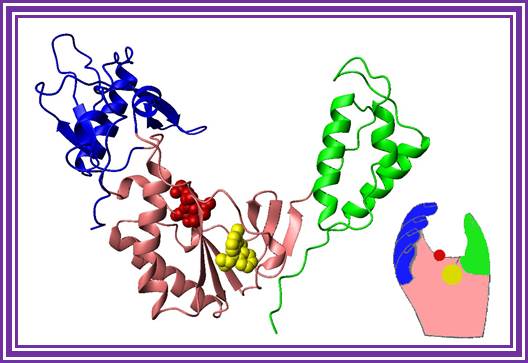
A Handy Enzyme
This ribbon representation of the RT active
domain illustrates its hand-like structure, showing fingers (blue), palm (pink)
and thumb (green). The active site (red atoms), where DNA is elongated, is in
the palm region. Also shown is an RT-inhibitor drug (yellow) in the pocket
where it binds.
http://www.psc.edu/
This is extensively used during second strand synthesis of cDNA. It requires RNA or DNA primers of 8 or more nucleotide long. These enzymes show low processivity and low fidelity. In most of the labs this enzyme is used for the preparation of first strand of cDNA. Its high temperature dependence of 42oC is helpful in removing ant secondary structures of the template. But Melony Murine leukemia enzyme is single stranded and it is mostly used in cDNA preparations.
Poly-(A) Polymerase.
This enzyme is a tetramer 33KD each. It has unique property of adding adenine nucleotides to 3’ end of any RNA. Only few nucleotides are required at the 3’ end of the RNA.

http://www.ebi.ac.uk/
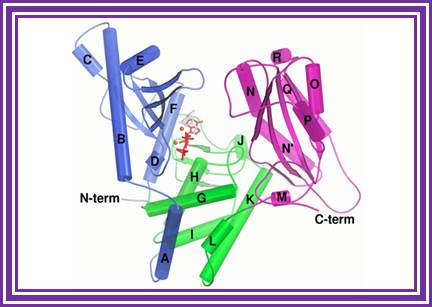
Some of the viral RNAs don’t have poly (A) tail; in such situations one can add poly (A) tail, which is of great help in preparation of ds CDNA from them.
Polynucleotide Phosphorylase:
It is tetramer, 60-70KD each. It can add rNDPs to 3’ ends of RNA and require Mn2^+. This can polymerize free nucleotides to generate polynucleotide chains. It can also act as nuclease.
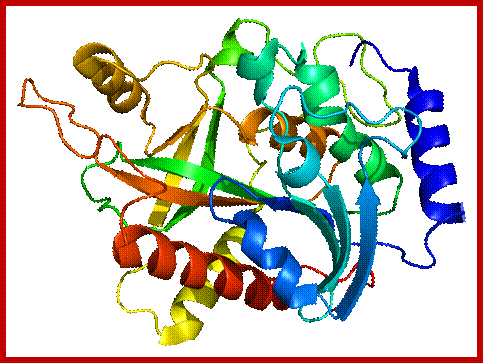
https://en.wikipedia.org/
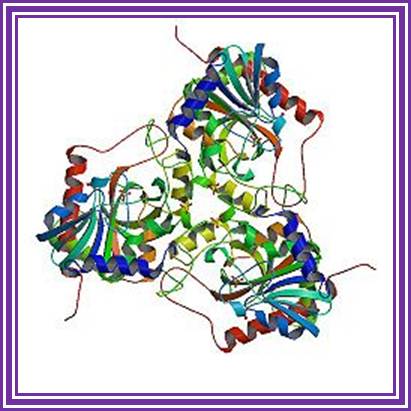
Polynucleoside Phosphorylase: http://www.creativebiomart.net/
Terminal Deoxynucleotidyl Transferase:
It is a dimmer consists of 26.5 and 8KD proteins. It adds dNTPs to the 3’ OH ends of ss DNA or dsDNA. It is template independent. This can be used for 3’ end labeling. This can also be used for generating 3’end tails of required nucleotides for cloning purpose.
General Structure of TdT. Note the "ring-like" shape of the enzyme. Figure taken from Delarue, et al. (2002) with permission. http://www.bio.davidson.edu/
T4 Polynucleotide Kinase:
It is obtained from T4 phages. It’s molecular mass is 33KD, it is a tetramer. It transfers gamma phosphate from the dATP to the 5’ of ds DNA or dsRNA. Even in the presence of 5’ ‘P’ group, it exchanges the gamma labeled phosphate, at the 5’ end. This is extensively used for 5’end labeling of DNA or RNA probes and the 5’ labeled DNA can be used for sequencing reactions.
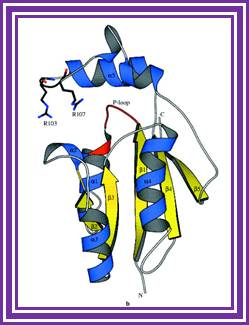
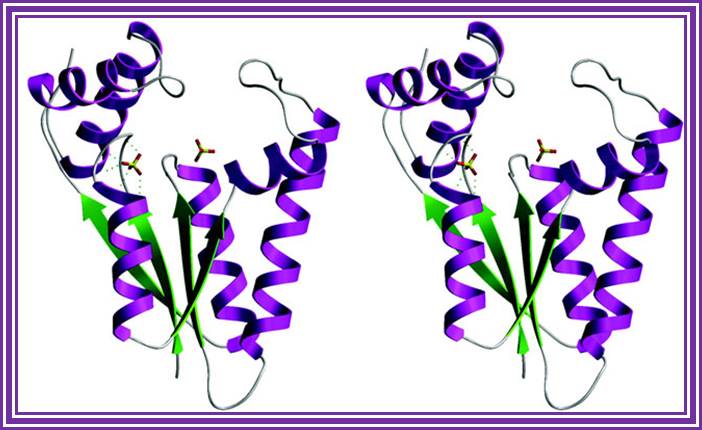
Structure an mechanism of T4 polynucleotide Kinase; an RNA repair enzymes; http://emboj.embopress.org/
RNA polymerases:
T7 RNA polymerase:
T7 RNA-pol (98-99KD), T3 RNA-Pol (100KD) and SP6 RNA-pol (96KD) can be used for synthesis of labeled RNA in large amounts provided if one uses their respective promoters in their constructs. T7 RNA Polymerase is a single-subunit enzyme produced by bacteriophage T7. It is highly specific for T7 promoter and terminator sequences. It has been widely used for the rapid synthesis in vitro of specific RNAs. These transcripts can be used directly as substrates for studies of RNA structure or metabolism. If the transcripts are suitably labeled, they can also be used as sensitive hybridization probes. T7 RNA Polymerase, along with chain-terminating nucleoside triphosphates. have been used for the direct sequencing of DNA. T7 RNA Polymerase is also used to generate capped mRNA for gene expression studies in oocytes and other cells. http://www.affymetrix.com/
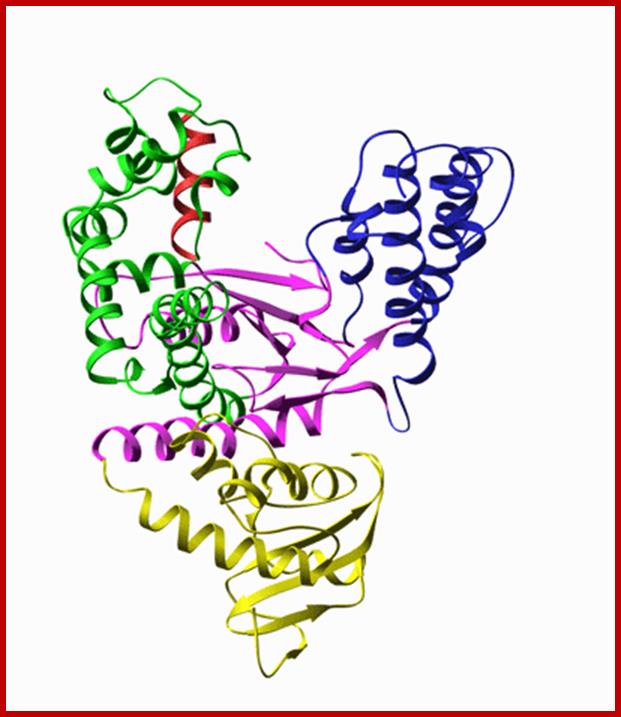
- It can be used as radiolabeled RNA probes, it can also be used as RNA vaccine, used as guide RNA for gene targeting, mRNA for in vitro translation and micro injection, RNA structure, processing and catalysis studies, RNA amplification and Anti-sense RNA for gene expression experiment ; https://www.neb.com/products
DNA Ligases:
There are different types of ligases of which E.coli DNA-ligase (74-77KD) and T4 DNA ligases (43KD) are extensively used. The former is NAD dependent and the later is ATP dependent. They are responsible for ligating the DNA ends. Double-strand DNA breaks arise pathologically as well as physiologically. There are two pathways for the repair of double-strand breaks (Liang et al. 1998). One is homologous recombination (HR), which occurs during late S and G2 of the cell cycle. The other pathway is nonhomologous DNA end joining (NHEJ). This is the predominant pathway during G0, G1, and early S phases of the cell cycle (Sonoda et al. 1998). Certain rare human diseases with autosomal recessive mode of inheritance are associated with a greatly increased cancer frequency which may reflect specific defects in DNA repair or replication. These disorders include Xeroderma pigmentosum, ataxia-telangiectasia, Fanconi's anemia and Bloom's syndrome and others, http://www.nature.com/
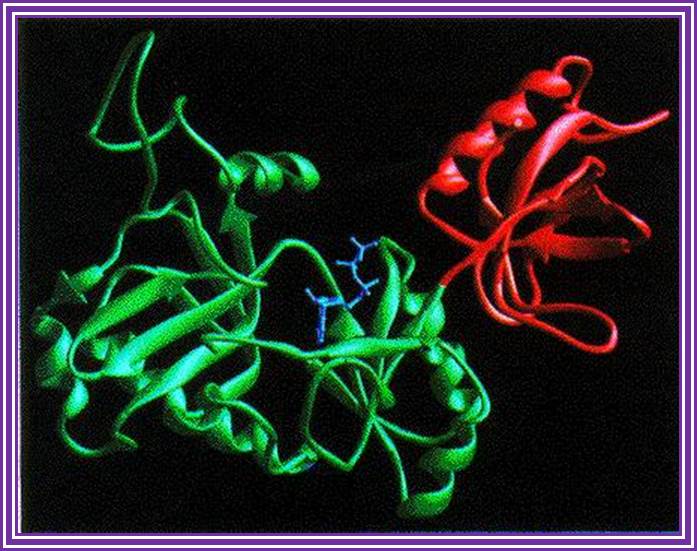
http://www.biochem.umd.edu/
Alkaline Phosphatase:
Alkaline Phosphotase enzymes are available from bacteria and calf intestine and many other sources. They are used to remove phosphate groups from 5’ ends of ssDNA or from dsDNA.
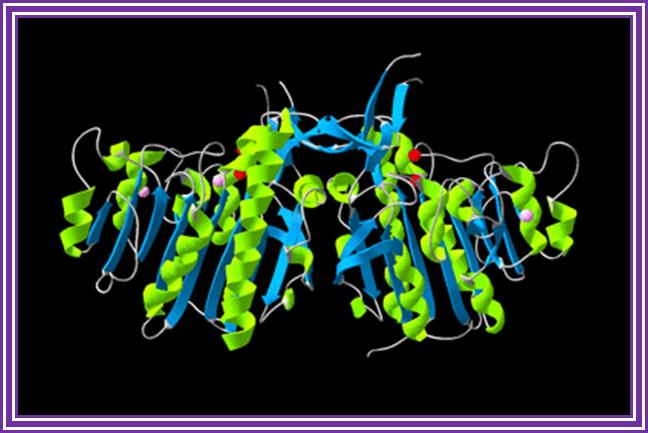
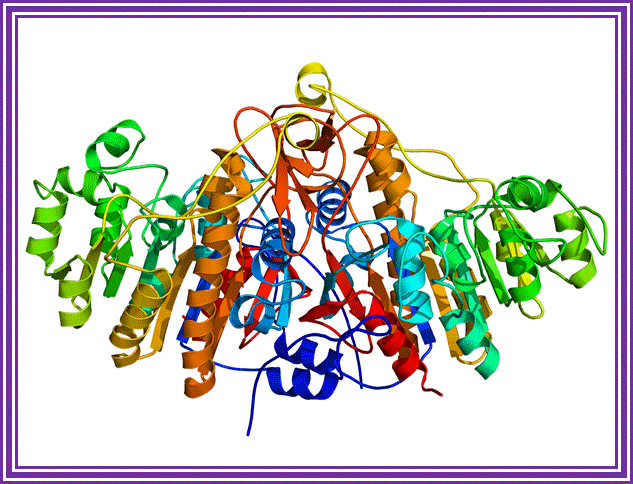
Alkaline Phosphatase; http://en.wikipedia.org/
Removal phosphate groups in vectors prevent self-ligation. However, one should be careful in using this enzyme for after treatment it has be removed completely by extracting the DNA 2 to 3 time in phenol-chloroform and then in chloroform once and precipitating in 95% ethanol in the presence of 0.15 molar sodium acetate. Alkaline Phosphatase from calf intestine can be easily inactivated by heat.
Guanylyl Transferase:
This enzyme used for capping of mRNA, that is lacking G cap. The enzyme transfers 7’CH3-GMP* from the 7’CH3-GTP to the 5 biphosphate or triphosphate by 5’ to 5’ phosphate bonding to generate 5’7’CH3 G-CH2-O-P*-O-P-O-P-O-CH2- ApXpXpX. The enzyme has a conserved lysine that serves as catalytic residue. The histidine tRNA in eukaryotes requires addition guanine residue before it is amino acetylated by Histidine tRNA synthetase?; it is specific to tRNA-His.
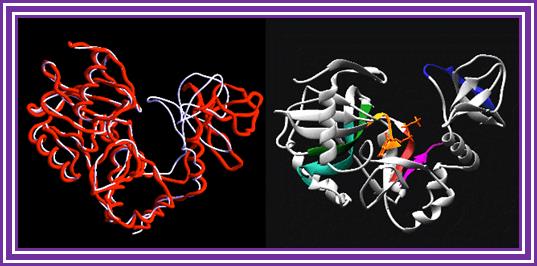

Guanylyl transferase: http://chemistry.umeche.maine.edu/
Most of the eukaryotic mRNAs contain ‘CAP; this s required for efficient Trnaslation of mRNA.
Nucleases:
Nucleases are a type of enzymes that cleave phosphodiester bonds between two nucleotides of nucleic acids. There are different kinds.
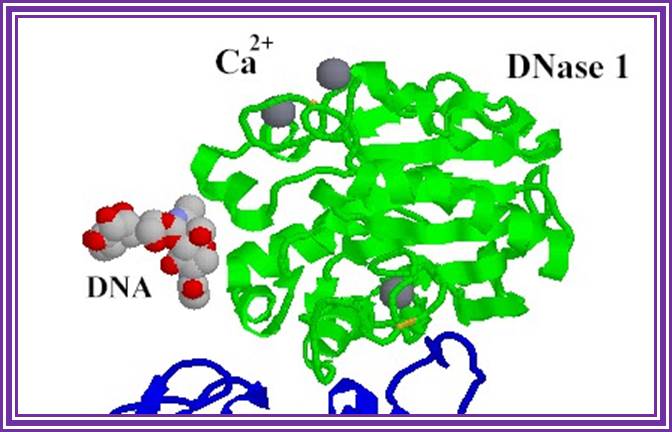
DNase I: RNase free:
Dnase I from bovine pancreas, 37KD, free from RNase is used for preparation RNA free of DNA. It is glycoprotein. It digests both ssDNA and dsDNA.
DNase 1;http://www.bms.ed.ac.uk/
This enzyme nicks double stranded RNA so it is used for labeling DNA by nick translation at very dilute concentration of enzyme. It is also used for DNase I foot printing and DNase I mapping.
Exonuclease III:
The enzyme is available as a recombinant product. Its Mol.wt is 28KD. This enzyme has the ability remove nucleotides after nucleotide from 3’ end of ds DNA either from blunt ends or from 3’ recessive ends. It also acts at nick region. Exonuclease is reported to have RNaseH activity. It also has proof reading activity.
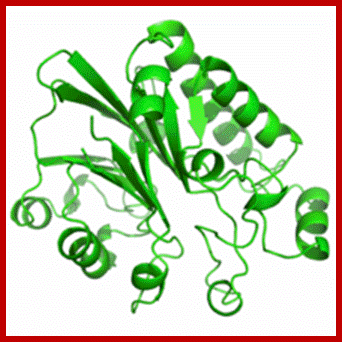
Exonuclease III crystal structure- from E.coli; https://en.wikipedia.org
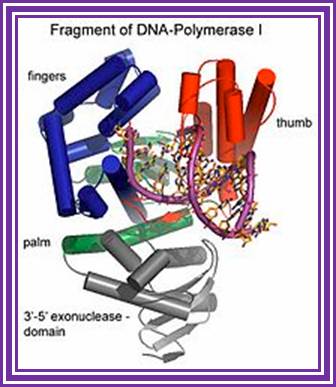
3’to 5’ exonuclease; http://en.wikipedia.org/
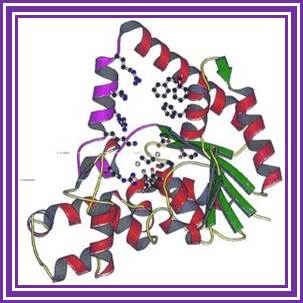
T5 -5’exonuclease
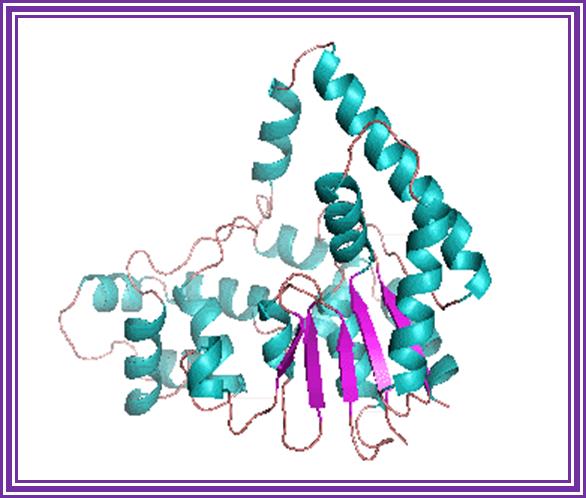
Flap Endonucleases, 5'-3' Exonucleases and 5' Nucleases; http://www.sayers.staff.shef.ac.uk/
T5 exonuclease degrades DNA from 5’ to 3’ direction, or it can initiate from 5’ when finds gaps; but this enzyme cannot act on supercoiled DNA. https://www.neb.com
It removes nucleotides after nucleotide to generate 5’P-mononucleotides. This enzyme is extensively used for deletional analysis of promoters to find out which part of the DNA segment can act as the promoter. If allowed for longer period it can generate single stranded DNA, but this enzyme does not act on single stranded DNA or RNA. This enzyme can also remove Apurinic and Apyrimidinic sites by endonuclease activity.
Lambda 5’ Exonuclease:
This ahs the ability remove nucleotides one after another from the 5’ end of the ds DNA either from the blunt end or 5’ over hangs. This is also useful in generating single stranded DNA and also for deletional analysis of the promoter elements.
S1 nuclease:
It is obtained from Aspergillus oryzae. It removes single stranded DNA or removes over hangs either from 3’ or 5’ end and produces a blunt ended DNA. Its Mol.wt is 38KD. It is a glycoprotein.

Aspergillus nuclease S (1); http://www.ebi.ac.uk/
Double stranded RNA and RNA: DNA hybrids are resistant to the action of this enzyme. This enzyme can be used S1 mapping of transcripts, one can remove 5’ or 3’ overhangs of DNA ends, and even they remove any hair pin structures.
Ribonuclease H:
The enzyme is obtained in large quantities from retroviral constructs.
RNase H; http://en.wikipedia.org/
The enzyme removes RNA by endonucleolytic cleavage of RNA from RNA: DNA hybrids. This enzyme used in the preparation of the second strand during ds cDNA synthesis.
Ribonuclease -A:
The main source is bovine pancreas. Its Mol.wt is 13.7 KD. In its pure form it is used for the preparation pure DNA free from RNA contamination in plasmid DNA preparation.
Ribonuclease A; http://en.wikipedia.org/
It hydrolyses 3’-5’ phophodiester bonds of RNA segments with 2’3’cyclic phosphates.
Proteinase K:
During preparation of nucleic acid from whole cells containing lot of proteins; to free from proteins this enzyme is used. This gives reasonable nucleic acid preparations.
Taq polymerase:
Reverse transcriptase:
DNA ligase; http://www.biochem.umd.edu/
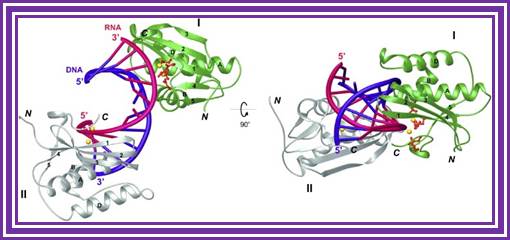
RNase H
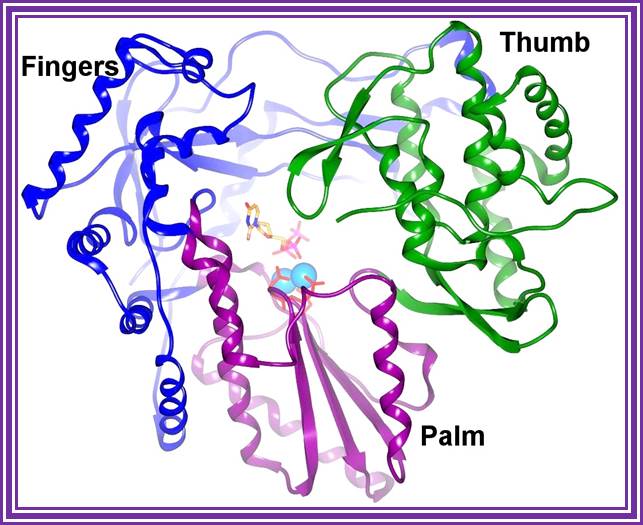
RNA polymerase?
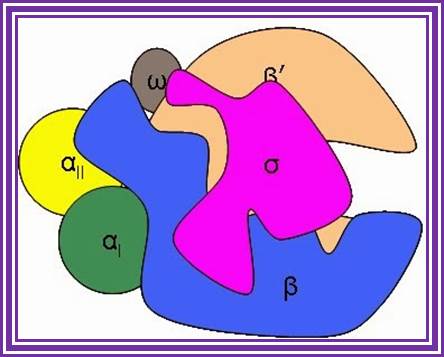
Bacterial RNA Polymerase; http://smallcollation.blogspot.com/

Fragment of DNA Pol I-3’-5’ exonuclease; DNA Pol I also contain 5’-3’ exonuclease activity http://en.wikipedia.org/
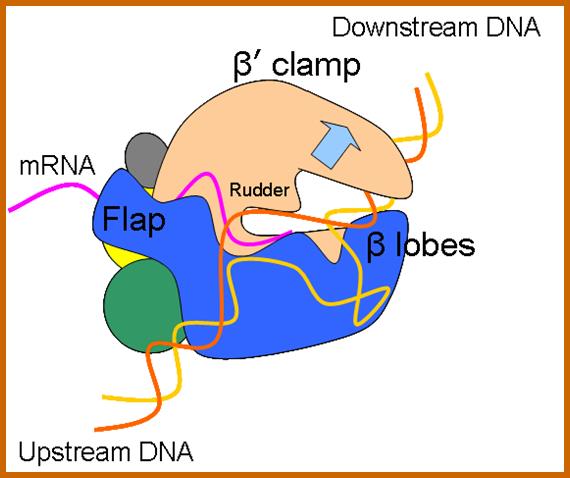
Bacterial RNAP
http://rationalwiki.org/
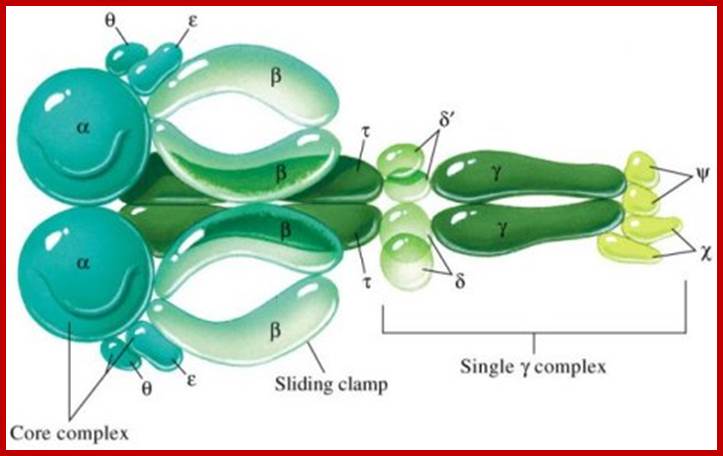 The
principal Polymerase of E. coli is Pol III, a multisubunit enzyme complex
The
principal Polymerase of E. coli is Pol III, a multisubunit enzyme complex
http://chemistry.umeche.maine.edu/
Severo Ochoa was awarded Nobel prize for the discovery of RNAP in 1969; in 2006 Roger Kornberg did detailed studies of RNA pol and its transcriptional process.
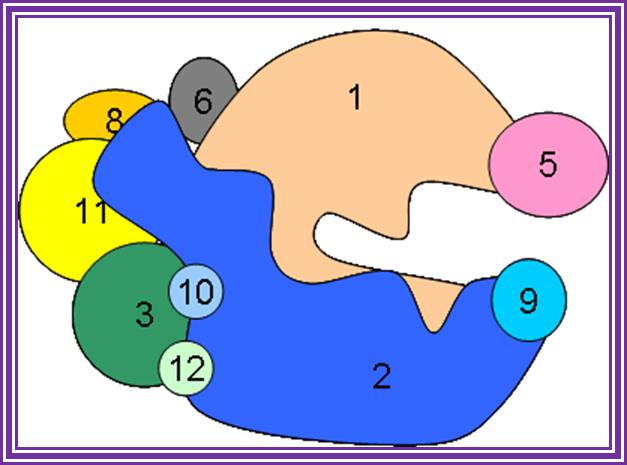
In eukaryotes RNA Pol 1 transcribes rRNAs.; http://www.conservapedia.com/
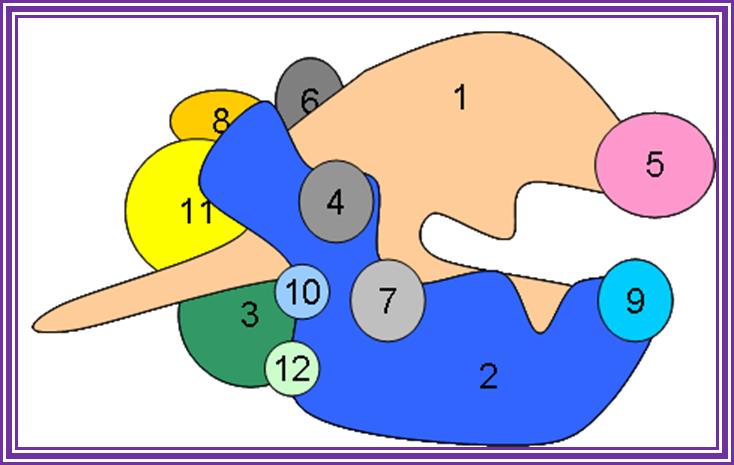
RNA Pol II produces mRNAs and snRNAs and micro RNAs.; http://www.conservapedia.com/
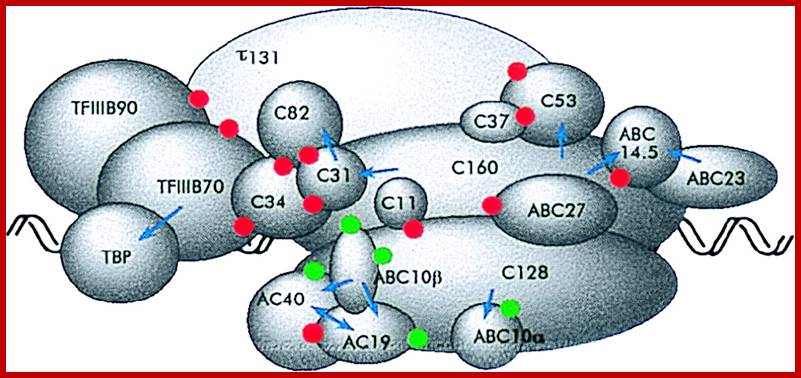
RNA pol III; http://www.ibb.unesp.br/
Proteinase K
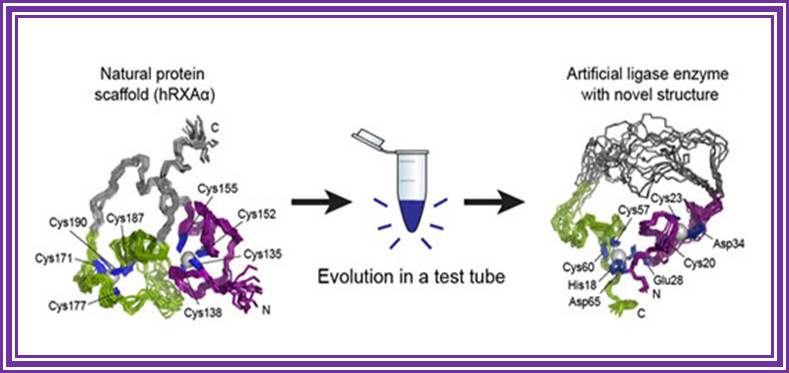
http://scitechdaily.com/